In Juan Carlos Izpisua Belmonte’s lab at the Salk Institute in La Jolla, California, the health of mutant mice suffering from kidney disease, muscular dystrophy, or diabetes improved after the scientists tweaked genes associated with the ailments. Belmonte and his colleagues, however, didn’t edit the genes themselves. Instead, they targeted the epigenome to modify the chemical and protein tags that sit on the chromatin and influence gene expression.
Scientists are developing molecular tools to edit these epigenetic tags, uncovering how they affect the noncoding regions of the genome and control gene expression. A better understanding of how the epigenome functions could lead to new, safer therapeutics for diseases, from cancer to elevated cholesterol. “There are a lot of really cool things you can do with epigenome editing that are really different than genome editing,” says Charles Gersbach, a biomedical engineer at Duke University.
Despite the revolutionary promise of genome editing, it has some drawbacks. For one, it changes a cell’s underlying DNA sequence, potentially creating unexpected effects that could cause cancer or other diseases. Epigenetic editing doesn’t change the DNA sequence, which can make it safer as a therapy, Gersbach says. Also, genome editing relies on DNA repair pathways, which insert new bases into the genetic code in an unpredictable fashion. Epigenome editing doesn’t cut DNA and doesn’t require stitching in bases to repair it, making the tools’ use across cell types more consistent, says Gersbach.
Engineering epigenome editors has its own challenges, however. For one, the biochemical diversity of epigenetic marks—histone proteins get methyl, acetyl, and other functional groups added to their tails, methyl groups on DNA itself take at least six different forms, and more types of marks are discovered each year—makes it more difficult to design just the right editor to target each tag individually without causing unwanted effects. Precisely targeting epigenetic marks at specific points on the genome is still difficult because the molecular mechanisms underlying epigenetic regulation are not well-understood, says Michael Elowitz, a genetic engineer at Caltech. “Right now, even though we have a lot of molecular knowledge, the underlying logic of the system is mostly mysterious.”
Here, researchers who have developed epigenome editing tools take The Scientist on a tour of their work.
EPIGENETIC OFF SWITCH
Researcher: Angelo Lombardo, geneticist, San Raffaele Telethon Institute for Gene Therapy
Project: Silencing gene expression for generations
Problem: For two decades, researchers have been using RNA interference, which disables messenger RNA post-transcription, or ZFNs and TALENs, artificial nucleases that disrupt the DNA sequence with targeted double-stranded breaks, to dial down or turn off the expression of a particular gene. But these methods provide only partial knockdown of a target gene or don’t target all of the alleles of a gene. Sometimes they don’t hit the right gene at all.
Solution: Eukaryotic cells naturally use epigenetics to repress gene expression. One well-understood mechanism for this process involves shutting off so-called endogenous retroviruses (bits of the genome derived from ancient retroviral infections) in stem cells of an embryo before it implants in the uterus. To turn off the leftover viral elements, cells use two specialized families of proteins, Krüppel-associated box containing zinc-finger proteins (KRAB-ZFP) and de novo DNA methyltransferases (DNMT). The former binds to viral sequences and removes methyl and acetyl groups from histone marks. The latter one adds methyl groups, locking in gene repression for future cell generations.
Lombardo and his colleagues tapped into this mechanism, repurposing one protein from each of the two protein families (they used KRAB-ZFP/KAP1 and DNMT3A) to shut off genes in differentiated human cells, such as T cells, rather than in stem cells. The researchers paired the two proteins with sequences that guided them to do their magic on epigenetic marks at three specific genes, effectively turning the genes off (Cell, 167:219–32.E14, 2016). Because each protein targets slightly different epigenetic marks, the two proteins edited the epigenome in different ways. One repressed gene expression rapidly, robustly, and reversibly, and the effect could be erased after a few rounds of cell division. The other also turned off the three genes, and the inhibition was lasting, fading out only after a few hundred cell generations. The fact that the repressive marks were stable a year later, after continuous cell division, is really impressive, says Gersbach, who was not involved in the study.
DISEASE DISCOVERY
Researcher: Charles Gersbach, biomedical engineer, Duke University
Project: Mapping cancer’s noncoding DNA with modified gene-editing tools
Problem: Millions of regions in the human genome don’t encode proteins. Some of those noncoding regions carry sequence variants associated with leukemia, breast cancer, and other complex diseases, but it’s unclear how these genomic segments regulate gene expression. Researchers have used CRISPR-Cas9 genome editing to interfere with certain genes and regulatory elements to explore their effects on health, but that technique does not work for all gene variants, nor does it correctly modify the regulatory elements that control them. What’s more, Gersbach notes, the technique can only turn genes off, not on, and it still relies on inserting or deleting DNA in the genetic code.
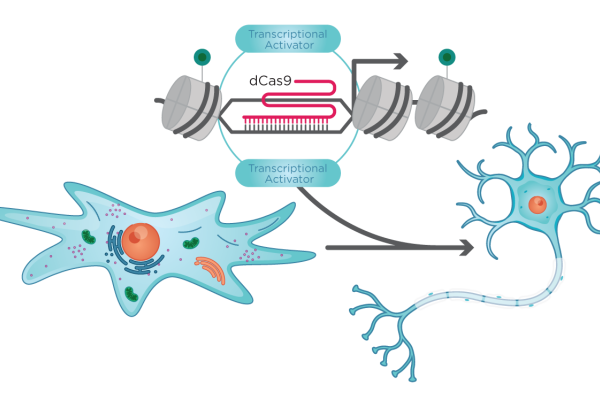
Solution: Gersbach and his colleagues designed a CRISPR-Cas9–based epigenome editor in which Cas9, mutated so as not to cut DNA, is fused to protein segments that modify the epigenome. Those protein segments home in on noncoding DNA regions known to act as promoters or enhancers, and recruit
epigenetic elements that either induce or repress DNA transcription.
The team used the editor in human cells to block noncoding regions that promote the transcription of genes in the b-globin locus, which has been implicated in leukemia, and the HER2 locus, which appears to play a role in breast cancer. The results, from experiments done in cancer cells and embryonic kidney cells, verified that previously known regulatory regions control gene expression at those loci. Targeting all candidate gene regulatory elements with guide RNAs and then comparing their abundances at different locations in the genomes of cells that expressed high or low levels of HER2 also revealed previously unidentified noncoding regions that regulate HER2 expression. Using the editor to turn on such noncoding regions provided an even more surprising result: activity in noncoding regions is specific to certain cell types, such as cancer cells or cells from specific organs—and therefore, to specific diseases (Nat Biotechnol, 35:561–68, 2017). “We’re essentially using epigenome editing to go in and map the noncoding genome,” Gersbach says.
EDITS FOR TREATMENT
Researcher: Juan Carlos Izpisua Belmonte, cell biologist, Salk Institute for Biological Studies
Project: Targeting gene expression to treat disease
Problem: Efforts to modify the CRISPR-Cas9 system to target specific methylation marks and histone modifications without cutting DNA have been relatively successful in cell culture, but translating the work to live animals has proven challenging. That’s partly because the CRISPR-Cas9 system can be too big to get inside cell nuclei, where epigenetic marks are edited.
Solution: Belmonte and his team set out to create a smaller epigenetic editor. They had a radical idea: Why not split the editor in two? One part, carried by an adeno-associated virus, would contain the Cas9 enzyme, and the other, carried by a second adeno-associated virus, would ferry the guide RNAs and gene switches. The two parts would work together to turn gene expression up or down.
The team used the tool to upregulate three genes—Klotho to treat kidney disease, Utrophin for muscular dystrophy, and Pdx1, which produces insulin, for diabetes—in live mice (Cell, 171:1495–1507.E15, 2017). Upregulating these genes compensated for disease-associated genetic mutations, improving symptoms in the mouse models of those diseases. The researchers also used the tool to overexpress certain long noncoding RNAs and genes rich in guanine and cytosine, which are challenging to sequence, revealing their biological functions. “Our paper represents a proof of principle that it is possible to treat diseases using an epigenetics approach,” Belmonte says.
CONTROLLING CHOLESTEROL
Researcher: Charles Gersbach, biomedical engineer, Duke University
Project: Blocking gene expression to lower cholesterol
Problem: The gene PCSK9 encodes a protein that regulates, and can sometimes prevent removal of, LDL (bad) cholesterol in the blood. An antibody drug that inhibits this protein to boost LDL removal has already been approved for treating familial hypercholesterolemia, a genetic disease in which the PCSK9 protein fails to remove LDL from the blood. Other therapies in development aim to inhibit the gene using synthetic oligonucleotides or gene editing. Because so much work has been done on PCSK9 and its connection with cholesterol, scientists know a lot about its expression. That foundation is helpful for building an epigenome editor that modulates it, says Gersbach. Using a standard tool based on the CRISPR-Cas9 system, Gersbach and his colleagues were easily able to turn off Pcsk9 by targeting specific epigenetic marks in cultured cells. But, as Belmonte’s team experienced, getting the tool to work in animal models was more difficult.
Solution: When using the CRISPR-Cas9 system, researchers typically take the bacterial immune system from a species such as Streptococcus pyogenes. But the epigenetic editing tool doesn’t always fit in the standard adeno-associated virus delivery system. So the team used the Cas9 system from Staphylococcus aureus; the gene-editing software is encoded by a collection of genes a kilobase smaller than that of S. pyogenes, and fits more easily in the delivery system along with guide RNAs and other molecules to modify gene expression.
As Belmonte’s team did, Gersbach and his colleagues created two delivery systems to transport the epigenome editor to the right region of the genome in the cells of living mice. One system carried a non–DNA-cutting version of Cas9 from S. aureus and protein segments that edit epigenetic marks, and the other contained guide RNAs that targeted the Pcsk9 gene. When the team injected the two-pronged system into adult mice and tested their liver cells, the mice had lower LDL cholesterol levels than animals that received a sham treatment (Nat Commun, 9:1674, 2018). One potential benefit of this method compared with oligonucleotides or gene editing, Gersbach says, is that it does not involve steps—such as cutting DNA or introducing foreign peptides—that could minimize efficiency or spur an unwanted immune response.
Originally published on The Scientist on January 1, 2019. View original story