The Department of Biostatistics and Bioinformatics (B&B) at Duke University is pleased to accept applications for its 2022 summer bioinformatics short-course in Microbiome Immunology Cancer (MIC). This two-week course will be held during the period of 06/06/2022 - 06/17/2022. This is the second year of a five-year training program funded by the National Cancer Institute (NCI). The course will be held in-person on the Duke School of Medicine campus in Durham, North Carolina if federal, state, local, and University guidelines at the time allow. If conditions make it necessary, the course will be transitioned to an online format.
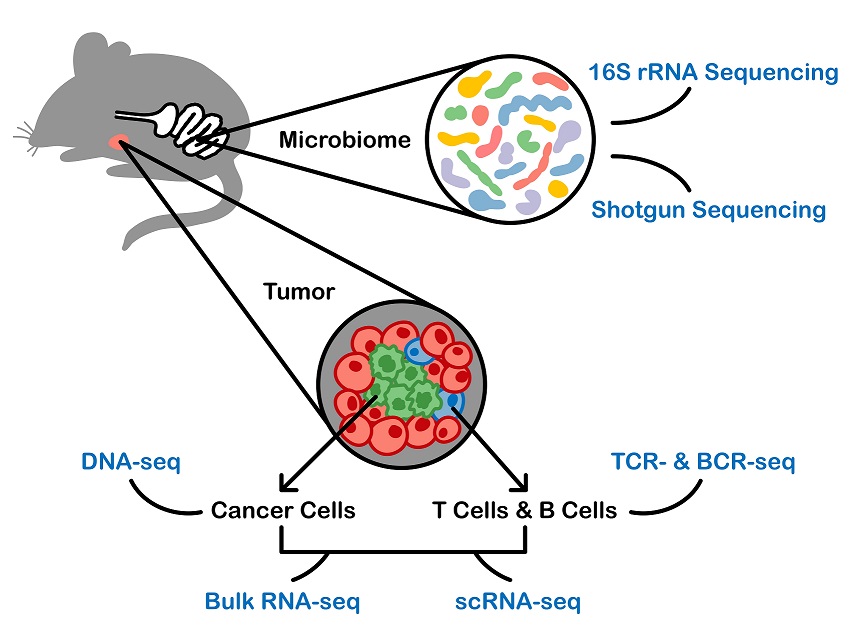
Modern cancer immunology research studies the complex interactions among the tumor, host immune system and microbiome, and depends on an array of technologies that were unimaginable a decade ago. To design, execute, and interpret experiments using these technologies, it is not sufficient for cancer researchers to only understand the biology of cancer; they must also understand the assay technology and the analysis methods used to interpret the resulting data. This complexity presents a challenging task and a huge unmet need in the education of cancer researchers, as what needs to be learned cuts across multiple disciplines: cancer biology, immunology, microbiology, statistics and bioinformatics. To address this pressing knowledge gap for cancer researchers, our summer course integrates the analysis of microbiome, immunology and cancer high-throughput data. This course will cover the biology, assays, online resources, bioinformatics pipelines and statistical methods needed to understand the analysis and interpretation of the results.
The first week of the course will consist of a preparatory sequence, and the second week will focus on an assay-specific sequence. The preparatory sequence will be devoted to teaching the fundamental elements and principles of the microbiome, cancer immunology, computing and statistics needed to prepare students to engage in bioinformatics analysis of high-throughput data. The week-two, assay-specific sequence will be devoted to teaching the requisite methodology and tools for conducting in depth analysis, from start to finish, of single-cell RNA-seq data. Subsequent offerings of the course will be devoted to other sequencing assays (e.g., 16s microbiome sequencing, TCR repertoire diversity analysis). Course participants are encouraged, but not required, to attend the course over multiple years.
The courses will teach best practices in reproducible analysis and provide hands-on practice needed to master performing the requisite analyses. The courses will be self-contained, in that no specialized background in biology, statistics or bioinformatics is assumed, although participants are expected to be sufficiently motivated to learn challenging interdisciplinary material.
There are no fees for attending the course since it is funded by a grant from the NCI. Because of this funding, accepted participants will be asked to formally affirm their commitment to attend the course in its entirety.
To apply: Applications for the course are currently being accepted on a rolling basis. Register here.
For additional information, please see the following:
If you have any remaining questions please contact the program administrator by email at miccourse@duke.edu .