Innovation
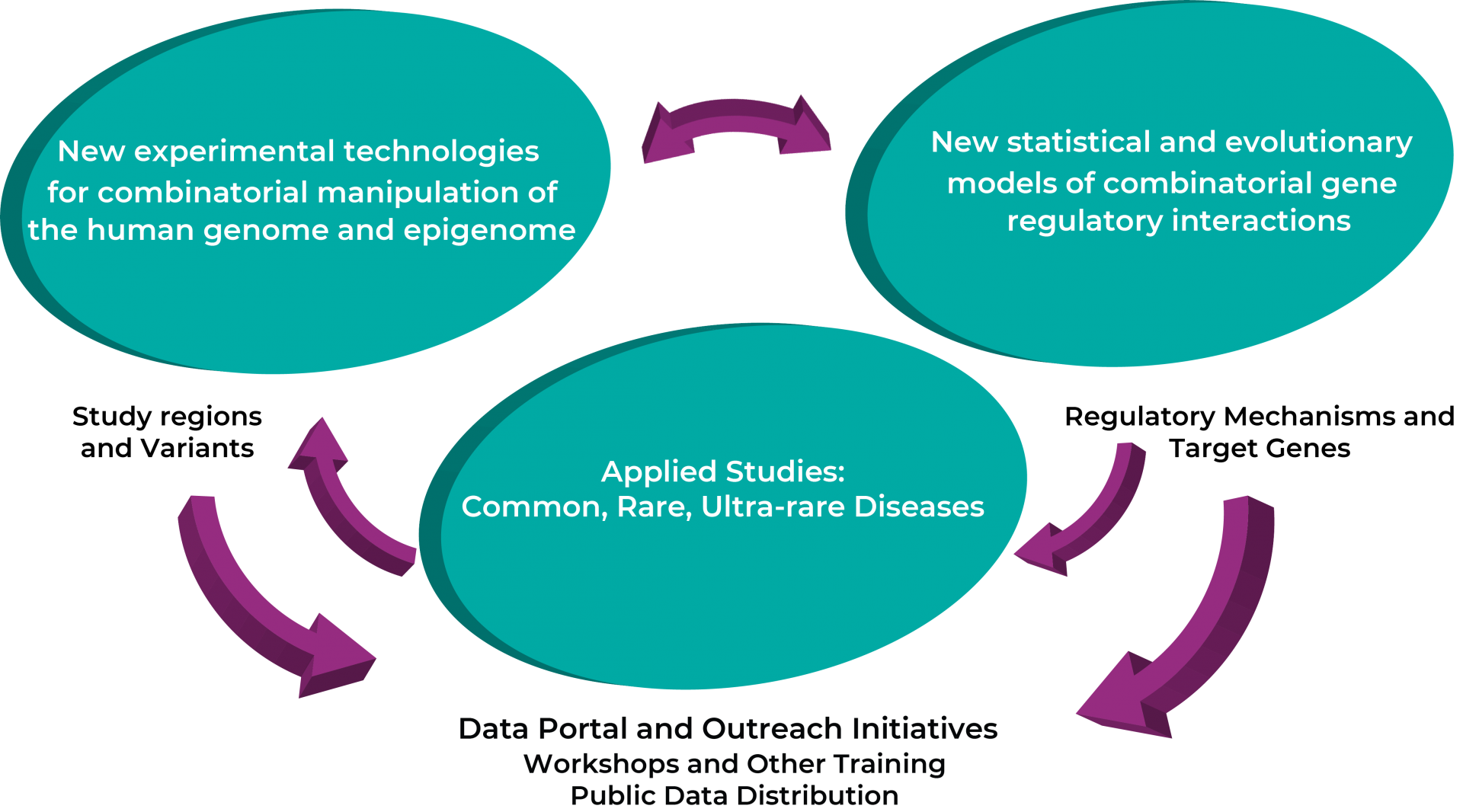
To achieve the goals of this project, we will drive synergistic innovation in four key areas:
- Experimental techniques to measure the function of combinations of cis-acting regulatory variants at several genomic scales;
- Statistical methods and evolutionary principles to model how those interactions influence gene expression and other phenotypes;
- Disease applications revealing different ways that combinations of regulatory elements and variants impact the spectrum from common to ultra rare disease; and
- The first public resource to disseminate datasets detailing the function of noncoding variants, alone and in combination, to basic scientists and translational scientists alike.
Those innovations together will be a windfall for truly understanding the many roles that gene regulation plays in human biology.
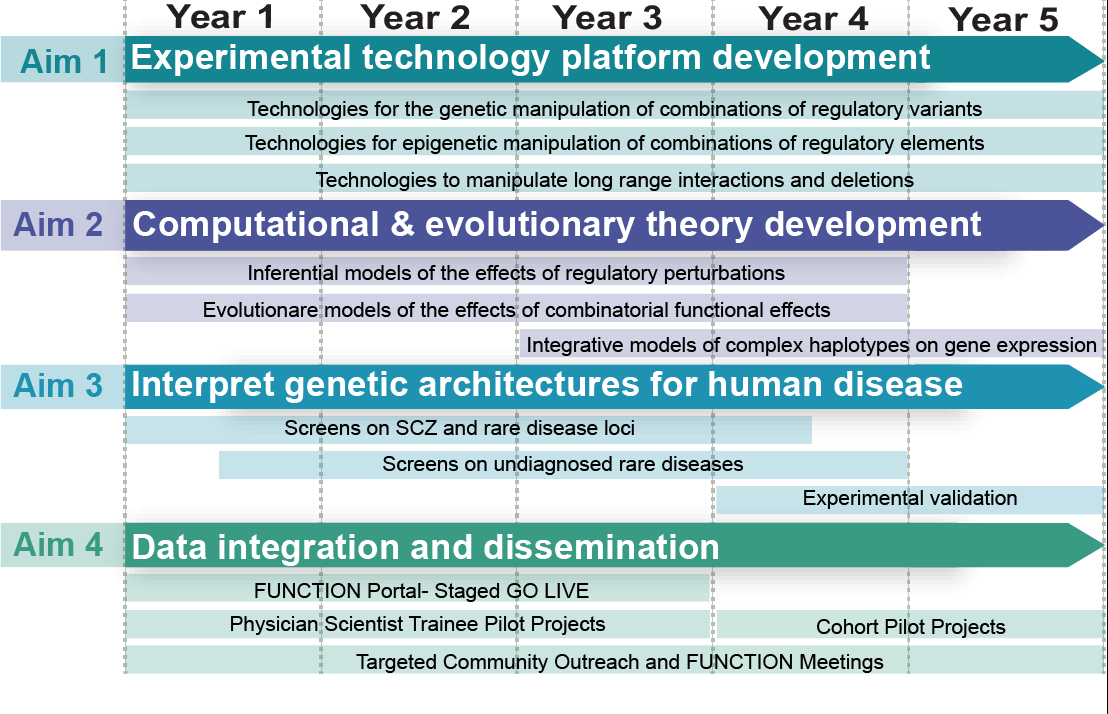
Aims
The Center for Gene Regulation (CCGR) will leverage a highly collaborative and interconnected study design to realize our goals. This project includes four aims:
New technologies to measure the combinatorial effects of regulatory elements and variants
Regulatory variants interact at different scales to impact human biology. Here, we will leverage our functional genomics expertise to develop the experimental foundation for measuring how combinations of noncoding variants at different scales contribute to an individual’s gene regulatory state. We will develop multimodal technologies for measuring combinatorial effects (i) within individual regulatory elements, (ii) between regulatory elements, and (iii) across megabase-scale regions of the human genome.
Statistical and evolutionary methods to estimate combinatorial regulatory effects
We will create an entirely new statistical and analytical framework to translate empirical measurements of the effects of combinations of noncoding variants into mechanistic and diagnostic interpretations and predictions. That will include new methods to infer and predict changes in regulatory activity and gene expression conferred by all of the noncoding variants that influence a gene in an individual, and new evolutionary theories to model selection and conservation based on noncoding gene regulatory function. The outcome will be the analytical foundation for interpreting the panoply of noncoding variants in whole genome sequences.
Identify combinatorial regulatory mechanisms of common, rare, and ultra-rare diseases
We have new evidence that regulatory variation contributes not only to common disease, but also rare and ultra-rare diseases. We hypothesize that different genetic structures of noncoding variation act at those extremes. We therefore will deploy the CCGR platform to evaluate the role of noncoding genetic interactions across the spectrum from common to ultra rare disease. Doing so will integrate basic and clinical expertise, and lead to new diagnoses and general principles for interpreting patient genomes.
The first community portal for the functional effects of regulatory elements and variants
We will establish the first online portal to collect and communicate measures of the regulatory activity of the noncoding genome. The portal will integrate results from the CCGR experiments and analyses with those from external studies by both consortia and individual labs. In doing so, we will create a key resource for researchers to incorporate assays of regulatory element activity in their own research. To enhance community impact, we will also implement training and outreach initiatives on the use of the portal and related CCGR technologies.